Introduction
Atrial fibrillation (AF) is the most common form of cardiac arrhythmia in clinical practice and causes significant morbidity and mortality [1]. Although many risk factors for AF have been identified, including advanced age, hypertension, structural heart disease, and congestive heart failure, a subset of younger subjects develop AF in the absence of established risk factors. The mechanisms underlying AF are not fully understood, but growing evidence supports an underlying heritable component and that genetic variation may play a role in the pathogenesis of AF [2]. Familial cases among young patients with lone AF underscore a genetic basis for the disease, and research implicates pathogenic mutations and risk-conferring functional polymorphisms in AF development. Recently, genome-wide association studies (GWAS) performed in populations of European-descent have identified several genetic variants associated with AF [3]. To date, three genomic regions / candidate genes have shown strong association with AF: 4q25, near paired-like homeodomain transcription factor 2 (PITX2); zinc finger homeobox 3 (ZFHX3) on 16q22; and potassium intermediate/small conductance calcium-activated channel, subfamily N, member 3 (KCNN3) on 1q21 [4,5]. The 4q25 genetic locus is adjacent to PITX2, a transcription factor that determines left-right asymmetry of the heart [6]. Recently, non-coding single nucleotide polymorphisms (SNPs) on chromosome 4q25 have been associated with the occurrence of AF in different populations [4,7]. The sequence variant rs2106261 is an intronic SNP located in the ZFHX3 gene, also called AT motif-binding factor 1 (ATBF1), on chromosome 16q22, and was highly associated with AF in a Chinese GeneID study [8]. The SNP in KCNN3, on chromosome 1q21, was most significantly associated with lone AF in people of European ancestry in a study of 1,335 individuals with lone AF and 12,844 unrelated individuals without AF [9]. One of the long-term> risk factors for AF is prolonged PR interval on electrocardiography (ECG) [10]. Increased P-wave duration is indicative of abnormal phenotypic characteristics such as left or right atrial hypertrophy and or enlargement, which reflect delayed intra- or inter-atrial conduction [11]. Multiple reports have also shown the value of P-wave prolongation as a predictor of AF after cardiac interventions such as coronary artery bypass surgery [12]. Genetic variants at 4q25 and 16q22, and PR intervals, have also been reported to predispose individuals to AF [13]. Recently, AF has been most effectively treated with radiofrequency catheter ablation (RFCA), a procedure that isolates and terminates the source of ectopic activity [14]. The use of RFCA is increasing worldwide. Given the cost and potential complications of this procedure, analysis of pre-procedural characteristics that stratify the procedural efficacy and safety might be helpful. Interestingly, the 4q25 SNP has been recently reported to predict the recurrence of AF after RFCA [15]. Here, we hypothesized that the phenotypes of AF patients, including electroanatomical characteristics of the left atrium (LA) and recurrence after RFCA, were modulated by the common AF susceptibility alleles, rs2200733 and rs6843082 at 4q25, rs2106261 at ZFHX3, and rs13376333 at KCNN3.
Materials and Methods
Study Population
The study protocol was approved by the Institutional Review Board of the Yonsei University Health System and adhered to the Declaration of Helsinki. All patients provided written informed consent. Participants included 659 Korean patients with AF who underwent RFCA and 659 age and sex-matched healthy controls who were included in the Ansan-Ansung cohort study in the Korean National Institute of Health (KNIH) database. Exclusion criteria were: 1) permanent AF refractory to electrical cardioversion; 2) LA anteroposterior diameter>55 mm as measured by echocardiogram; 3) AF with rheumatic valvular disease; 4) associated significant structural heart disease; and 5) prior AF ablation or maze surgery. Three-dimensional (3D) spiral computed tomography (CT) scans (64 Channel, Light Speed Volume CT, Philips, Brilliance 63, the Netherlands) were performed to visually define the pulmonary vein (PV) and LA anatomy. Trans-thoracic echocardiography (Sonos 5500, Philips Medical System, Andover, MA, USA or Vivid 7, GE Vingmed Ultrasound, Horten, Norway) was performed to evaluate the presence of structural heart disease, degree of LA remodeling, or ventricular function. Using trans-esophageal echocardiography, we ensured that no LA thrombus was present. All antiarrhythmic drugs were discontinued for at least five half-lives. Amiodarone was discontinued at least four weeks prior to the procedure. Anticoagulation therapy was maintained before RFCA.
Electrophysiologic Mapping and Radiofrequency (RF) Catheter Ablation
Intracardiac electrograms were recorded using the Prucka CardioLab™ electrophysiology system (General Electric Health Care System Inc., Milwaukee, WI, USA). Catheter ablation was performed in all patients using 3D electroanatomical mapping (St. Jude Medical Inc., Minnetonka, MN, USA) merged with 3D spiral CT. If the initial recording identified a sinus rhythm, an LA 3D electroanatomical voltage map was generated during atrial pacing by obtaining contact bipolar electrograms from 350–400 points on the LA endocardium. The bipolar electrograms were filtered from 32–300 Hz. For patients with the initial rhythm of AF during the procedure, we generated voltage maps during pacing following internal electrical cardioversion (2 to 10 J, biphasic shocks with R wave synchronization, anodal decapolar catheter in high right atrium to cathodal duo-decapolar catheter inside of the coronary sinus, Lifepak12, Physiocontrol Ltd.). We did not obtain an LA voltage map if the frequently re-initiating AF required electrical cardioversion more than three times. We generally performed RFCA in sinus rhythm after electrical cardioversion. However, if sinus rhythm could not be maintained due to immediate recurrence of AF, RFCA was performed while in AF. We used an open irrigated-tip catheter (Celsius, Johnson & Johnson Inc.; Diamond Bar, CA, USA; irrigation flow rate 20 to 30 mL/min; 30 W; 47°C) to deliver radiofrequency energy for ablation (Stockert generator, Biosense Webster Inc.; Diamond Bar, CA, USA). All patients initially underwent circumferential PV isolation and bi-directional block of the cavo-tricuspid isthmus. For patients with paroxysmal AF, additional linear ablation or complex fractionated atrial electrogram (CFAE) ablation, guided by the 3D-CFAE-cycle length map, was conducted at the operator's discretion. In patients with persistent AF, circumferential PV isolation, cavo-tricuspid isthmus block, roof line, posterior inferior line, and anterior line were performed as a standard lesion set. If AF persisted, CFAE ablation was done depending on the operator's decision. If AF persisted after internal cardioversion, despite administration of the aforementioned ablation protocols, procedures were stopped. The procedural endpoint was defined as when there was no immediate recurrence of AF after cardioversion during isoproterenol infusion (5–20 μg/ min). If non-PV foci were identified under isoproterenol, they were ablated.
Post-ablation Follow-up
Patients were asked to visit an outpatient clinic one, three, six, and 12 months after RFCA and every six months thereafter for follow-up. ECG was performed at each visit and anytime the patient reported palpitations. A Holter ECG (24- or 48-hour) and/or event recorder was performed on patients at every six-month follow-up, and whenever the patient complained of symptoms, for at least two years in accordance with the 2012 ACC/AHA/ESC expert consensus guidelines for AF management [14]. We defined recurrence of AF as any episode of AF or atrial tachycardia of at least 30 seconds in duration. If AF was found on ECG during follow-up within three-months of RFCA, the patient was diagnosed with early recurrence of AF. Any AF recurrence thereafter was diagnosed as clinical recurrence and antiarrhythmic medications were prescribed.
Off-line Analyses of Color-coded 3D Maps and CT Images
Color-coded voltage maps were generated by recording bipolar electrograms and measuring peak-to-peak voltage during high right atrial pacing with a cycle length of 500 ms. We analyzed the color-coded voltage maps, that had been converted to image files, of both anterior-posterior (AP) and posterior-anterior (PA) views. PVs were not included in the analysis. Digital measurements of color-coded voltage maps were performed by a single investigator blinded to the clinical information of the maps, using a consistent method. The percentage of color-coded areas in each quadrant of the voltage maps was analyzed using customized software (Image Pro), and referenced to the color scale bars. Low voltage areas were coded gray and defined as LA voltage≤0.2 mV. High voltage areas were coded purple and defined as LA voltage>5.0 mV. The mean LA voltage was calculated by adding the area of each color (as a percentage), multiplying by the corresponding voltage, and dividing by the total area of LA. The reference distance was measured by the inter-electrode distances of coronary sinus catheters (duodecapolar catheter, St. Jude Medical Inc. Minnetonka, MN, USA.). To calculate local conduction velocity, the conduction distance was measured on the AP and PA views of the isochronal map, and the measured distance divided by the time difference. The 3D spiral CT images of LA were analyzed on an image processing workstation (Aquarius, Terarecon Inc., USA). Each LA image was divided into portions according to embryological origin as follows: the venous LA (posterior LA including the antrum and posterior wall), anterior LA (excluding the LA appendage and venous LA), and LA appendage. The absolute and relative volumes of each portion were calculated and compared.
Electrocardiography
Standard 12 lead ECGs were recorded for a few hours to a day before the catheter ablation procedure. Digitally stored ECGs were magnified on a high resolution computer screen and measured with a computerized caliper (Cardio Calipers, ver 3.3, ICONICO, NY, USA). Lead I, II, III, aVL, aVF, and Lead V1 were measured using Cardio Calipers. PR intervals were automatically measured with the GE Marquette MAC 5000 (GE Marquette Inc., Milwaukee, WI). P wave durations in Lead I, II, III, aVL, and aVF were measured from the onset of the P wave to the return to the baseline. P wave amplitudes were measured from the crest of the P wave to the trough of the wave. The biphasic deflection in Lead V1 was determined from the following: the duration of the initial positive deflection as measured from the onset of the P wave to the return to its baseline, before the onset of the negative deflection; the duration of negative P wave deflection as calculated by subtracting the duration of the positive deflection from the duration of the entire P wave; and the amplitude of the terminal negative deflection as measured from the trough of the wave to the baseline (Figure 1).
Genotyping
Whole blood samples were collected using EDTA as an anticoagulant and DNA was extracted for genetic analyses. Genotyping of 4q25 rs2200733, PITX2 rs6843082, ZFHX3 rs2106261, and KCNN3 rs13376333 was performed using validated TaqMan assays (Life Technologies, Carlsbad, CA, USA). PCR reaction mixtures contained 0.9 µm each of the forward and reverse primers, 0.2 µm each of the fluorescein amidite and VIC minor groove binder sequence-specific probes, 3 ng DNA, 5.0 mM MgCl2, and 1xTaqMan Universal PCR Master Mix containing AmpliTaq gold DNA polymerase in a 5.5 µL reaction volume. All SNPs had a call rate of >99%. All genotyping was performed by laboratory personnel blinded to all clinical data.
Statistical analysis
All values are expressed as the mean ± standard deviation. Multiple variables including clinical features, echocardiography parameters, and degree of electroanatomical remodeling of LA were compared according to different genotypes. The Student's t-test, Mann-Whitney U test, or Kruskal-Wallis H test was used, as appropriate, for comparisons between continuous data. The chi-square test or Fisher's exact test were used for comparisons between the categorical data. A chi-square test was used to test if the genotype distribution deviated from Hardy-Weinberg equilibrium. Kaplan–Meier survival analysis with a log rank test was used to present the recurrence of AF. Multivariate analysis was performed with the Cox stepwise forward regression model to determine the independent predictors of the recurrence of AF and odds ratios (ORs) using 95% confidence intervals (CIs). P-values less than 0.05 was considered statistically significant. All statistical analyses were performed using SPSS version 20.0 (SPSS Inc. Chicago, IL, USA).
Results
Clinical characteristics
All AF patients underwent RFCA, 457 of them had paroxysmal atrial fibrillation (PAF) and 202 had persistent AF (PeAF). The baseline characteristics of the AF patients and controls are summarized in Table 1.
Table 1.
Baseline characteristics of AF patients and control
Frequency of genotypes in Korean patients with AF vs. controls
Table 2 summarizes the frequency of the genotypes of the common AF susceptibility alleles. 4q25 rs2200733 TT genotype (51.5 versus 25.8%, p<0.001), CC genotype (7.7 versus 23.4%, p<0.001), PITX2 rs6843082 GG genotype (81.0 versus 49.8%, p<0.001), AA genotype (1.4 versus 8.8%, p<0.001), ZFHX3 rs2106261 AA genotype (24.5 versus 9.1%, p<0.001), and GG genotype (26.3 versus 49.5%, p<0.001) were significantly different between the patient and control group. In contrast, the distribution of rs13376333 genotype was not significantly different between patients and control.
Table 2.
Genotype distribution
Comparison of AF patients with and without the rs2200733 variant
There were no significant differences in clinical and electroanatomical parameters between the rs6843082 SNP variant and non-variant groups. The comparison of AF patients with the non-risk allele of rs2200733 (CC genotype) and risk allele (TT+CT genotype) is shown in Table 3. Patients with the risk allele (T) exhibited a lower rate of CHF (2.1 versus 8.0%, p=0.012). rs2200732 variant allele carriers had larger LA volume than non-variant carriers (128.5±40.7 versus 113.5±29.2 mL, p=0.005). When considering ECG parameters, the risk allele was associated with longer PR interval (185.9±32.6 versus 174.8±2 1.9 ms, p=0.018), longer duration of negative P terminal force in V1 (57.3±24.3 versus 48.8±23.9 ms, p=0.018), and higher amplitude of negative P terminal force in V1 (0.07±0.04 versus 0.05±0.04 mV, p=0.015). In addition, risk allele carriers tended to have a higher, but not statistically significant, incidence of AF recurrence.
Table 3.
Comparison of AF patients with and without the rs2200733 variant
Comparison of AF patients with and without the rs2106261 variant
The characteristics of AF patients, according to rs2106261 genotype, were compared (Table 4). The AA genotype carriers showed shorter P-wave duration in Lead aVL (93.5±22.5 versus 98.6±19.3 mV, p=0.026) and higher venous LA voltage (1.18±0.97 versus 0.94±0.71 mV, p=0.023) (Figure 2). In addition, clinical recurrence occurred more frequently in patients with the AG + GG genotype than the AA genotype (28 versus 20%, p=0.049).
Figure 2.
Representative images of color-coded 3D voltage map of LA. LA voltage, especially venous LA voltage, is higher in a patient with rs2106261 AA genotype (A) than AG+GG (B). LA, left atrium

Table 4.
Comparison of AF patients with and without the rs2106261 variant
Characteristics and predictors of clinical recurrence of AF after RFCA
Clinical recurrence of AF after RFCA was observed in 26.2% of patients. We compared the characteristics of AF patients with and without clinical recurrence of AF (Table 5). Patients with recurrence were more likely to have PeAF (46.3 versus 23.5%, p<0.001), larger LA (42.8±6.4 versus 41.1±6.2 mm, p=0.002), and longer PR interval (189.9±31.2 versus 183.4±32.7 ms, p=0.027). Procedure time (p<0.001), fluoroscopic time (p=0.003), and ablation time (p<0.001) were all significantly longer in patients with recurrence of AF. The ZFHX3 rs2106261 variant allele was more prevalent in patients with AF recurrence (81.5 versus 73.7%, p=0.049). We performed Kaplan-Meier survival analysis according to genotypes (Figure 3). During an 18.8±8.5 month follow-up period patients with the 4q25 rs2200733 risk allele (T) showed a trend of higher risk of recurrence (Log Rank p=0.061, Figure 3A). ZFHX3 rs2106261 variant allele carriers had significantly higher recurrence rate than non-variant (Log Rank p=0.015, Figure 3B). When patients were assigned to three groups according to the number of variant alleles (Group A: no variant, n=15; Group B: 1 variant, n=158; and Group C: 2 variants, n=439) there were no significant differences in age, gender, paroxysmal AF, CHADS2 (congestive heart failure, hypertension, age=75 years, diabetes mellitus, stroke) score, LA size, and medications. However, venous LA volume was significantly greater in patients with the variant allele (Group A versus B & C; 16.9 versus 22.3 and 22.9 mL/m2; p=0.023 & 0.020, respectively). Kaplan-Meier survival analysis showed incremental prognostic value according to the number of variant alleles (Figure 4, Log Rank p=0.015). Univariate Cox regression analysis identified that PeAF, LA size, and the number of variants in rs2200733 and rs2106261 were associated with the recurrence of AF after RFCA. Multivariate analysis showed that PeAF (OR 1.677, 95% CI 1.176–2.381, p=0.004) and the number of variants (OR 1.552, 95% CI 1.099–2.222, p=0.015) were independent predictors for AF recurrence (Table 6).
Figure 3.
(A) Kaplan-Meier curves of AF free survival in patients with and without the rs2200733 variant allele after RFCA of AF. (B) Kaplan-Meier curves of AF free survival in patients with and without the rs2106261 variant allele after RFCA of AF. AF, atrial fibrillation; RFCA, radiofrequency catheter ablation.
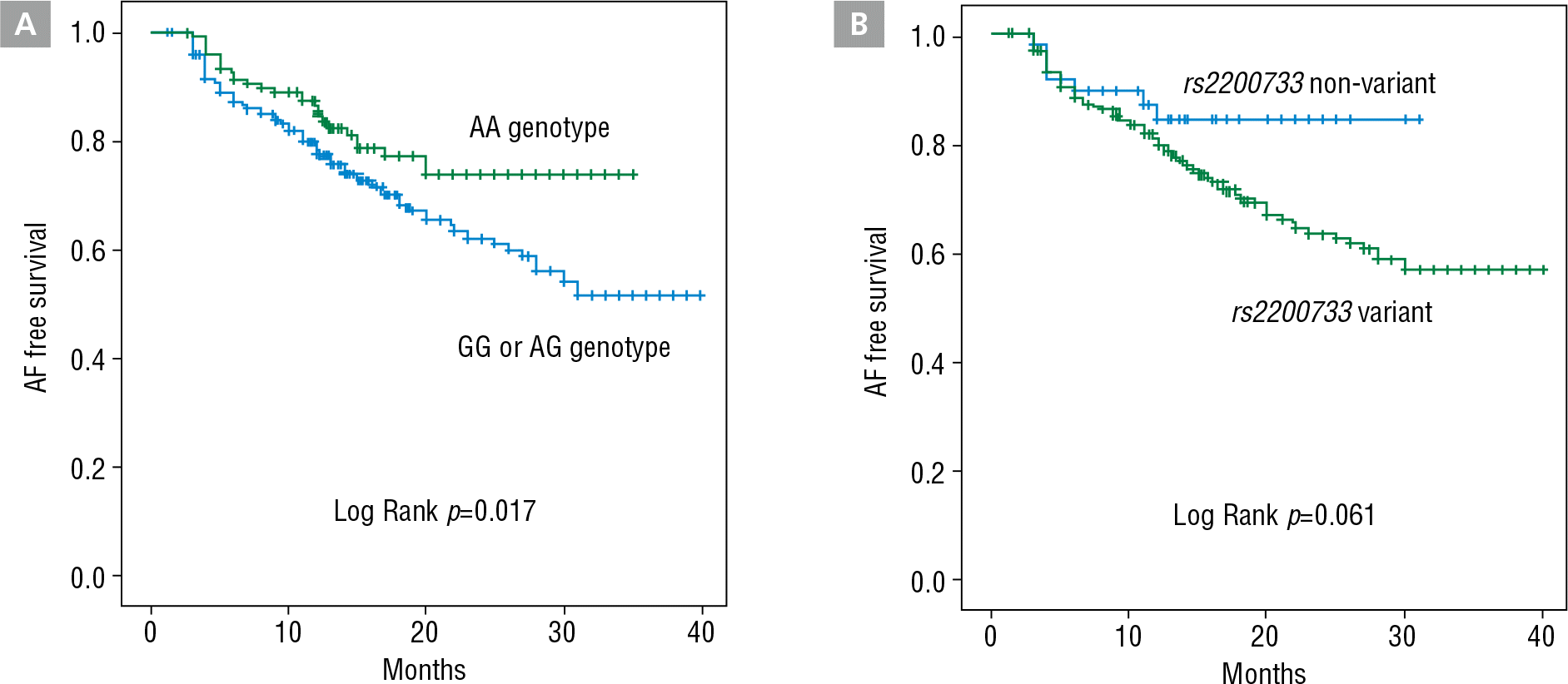
Figure 4.
Kaplan-Meier curves of AF free survival according to the number of variant alleles of rs2200733 and rs2106261 (Group A: no variant; n=15, Group B: 1 variant; n=158, Group C: 2 variants; n=439) AF, atrial fibrillation.
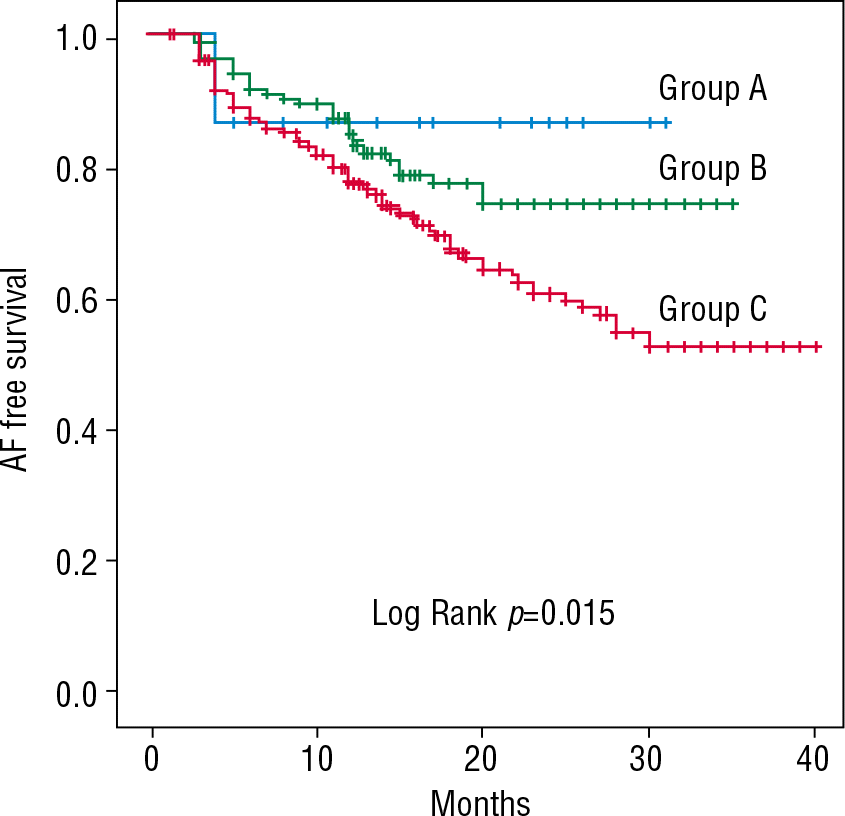
Table 5.
Comparison of AF patients with and without clinical recurrence of AF
| Recur (+) (n=165) | Recur (−) (n=486) | P value | |
| Age (years) | 58±11 | 57±11 | 0.312 |
| Male gender (%) | 78.0 | 74.8 | 0.402 |
| PAF (%) | 53.7 | 76.5 | <0.001 |
| BMI (kg/m2) | 24.9±3.0 | 24.8±2.6 | 0.749 |
| CHADS2 score | 0.87±0.99 | 0.84±1.03 | 0.714 |
| CHF (%) | 2.8 | 2.4 | 0.754 |
| Hypertension (%) | 45.2 | 45.7 | 0.918 |
| Age >75 (%) | 5.1 | 3.7 | 0.427 |
| DM (%) | 14.1 | 12.6 | 0.618 |
| Stroke/TIA (%) | 23.2 | 20.7 | 0.791 |
| ECG parameters | |||
| PR (ms) | 189.9±31.2 | 183.4±32.7 | 0.027 |
| P dur II (ms) | 124.2±21.4 | 121.7±19.9 | 0.173 |
| P dur V1 (+) (ms) | 60.4±17.9 | 58.6±22.2 | 0.326 |
| P dur V1 (−) (ms) | 57.1±25.2 | 56.6±24.2 | 0.813 |
| P dur V1 (+/−) (ms) | 116.9±24.2 | 115.1±21.4 | 0.394 |
| P amp I (mV) | 0.08±0.03 | 0.09±0.03 | 0.001 |
| Echo parameters | |||
| LA size (mm) | 42.8±6.4 | 41.1±6.2 | 0.002 |
| LVEF (%) | 62.9±8.1 | 63.9±7.8 | 0.158 |
| E/E' | 10.2±3.8 | 10.1±4.2 | 0.824 |
| LA volume on CT (mL) | |||
| LA | 72.1±21.5 | 70.0±22.3 | 0.329 |
| Venous LA | 22.6±8.4 | 22.3±8.4 | 0.714 |
| LA appendage | 5.9±2.7 | 5.7±2.9 | 0.480 |
| Anterior LA | 42.8±14.9 | 41.3±15.4 | 0.309 |
| LA voltage (mV) | |||
| Mean LA | 1.07±0.51 | 1.20±0.62 | 0.056 |
| Venous LA | 0.89±0.67 | 1.00±0.79 | 0.219 |
| LA appendage | 2.11±1.25 | 2.40±1.47 | 0.070 |
| Anterior LA Conduction velocity (m/sec) | |||
| Anterior | 0.55±0.25 | 0.57±0.30 | 0.727 |
| Posterior | 0.60±0.31 | 0.65±0.37 | 0.248 |
| Procedure | |||
| Procedure time (min) | 207.0±61.6 | 185.2±37.8 | <0.001 |
| Fluoroscopic time (min) | 52.5±21.7 | 45.7±27.1 | 0.003 |
| Ablation time (min) | 94.6±31.1 | 83.9±24.4 | <0.001 |
| Genotype distribution* | |||
| rs2200733 (4q25) | 95.5 | 91 | 0.067 |
| rs6843082 (PITX2) | 20.3 | 19.0 | 0.733 |
| rs2106261 (ZFHX3) | 81.5 | 73.7 | 0.049 |
Table 6.
Cox regression analysis of AF recurrence after catheter ablation
Discussion
Here, we found that polymorphisms 4q25 PITX2 and 16q22 ZFHX3, detected from peripheral blood, were associated with AF. The 4q25 rs2200733 and ZFHX3 rs2106261 genotypes were associated with electroanatomical remodeling of LA. The presence of the variant allele of ZFHX3 rs2106261 was also associated with AF recurrence after RFCA.
AF is a hereditable disease and related to genetic poly-morphisms
AF is known to be a degenerative disease with a multi-factorial background, such as aging and oxidative stress. Growing evidence shows that genetic variation may also play a role in the pathogenesis of AF [2]. Fox et al. [16] reported that a parental history of AF increased the risk of AF 1.9 times. Meanwhile, Lubitz et al.[17] showed a 39% increase in the incidence of AF in patients with a family history of AF in their 1st degree relatives. Recently, genome-wide association studies have revealed several nuclear polymorphisms significantly associated with AF. These include polymorphisms in genes such as KCNN3 on chromosome 1q21, PITX2 on chromosome 4q25, and ZFHX3 on chromosome 16q22 [4,5]. Common SNPs in a non-coding region of chromosome 4q25 have been shown to be associated with AF, and rs2200733 is the most reliable genetic variant associated with AF [3]. SNPs found in the 4q25 region are near PITX2, a gene believed to be involved in cardiac development, sinus node development, and susceptibility to atrial arrhythmias [18]. There is emerging evidence that highly conserved non-coding regions may act as regulatory elements and contribute to phenotypic diversity [19]. However, the mechanism by which genetic variation at the chromosome 4q25 locus leads to AF remains to be elucidated. Here, we evaluated the robust SNPs associated with AF: rs2200733 and rs6843082 at chromosome 4q25, rs2106261 at ZFHX3, and rs13376333 at KCNN3. The AF risk allele of SNPs rs2200733 and rs2106261 are much more frequent among Chinese subjects, in both affected and unaffected individuals[7,8] in concordance with our results. Contrary to the results of other studies, in our study population, rs13376333 of KCNN3 was not associated with AF [9]. These differences could be explained by intrinsic racial differences in myocardial membrane stability, myocardial conduction pathways, or genetic polymorphisms leading to different susceptibility to the development of AF. Another possibility is that the difference could be a result of complicated gene–gene and gene–environment interactions, which can dilute or accentuate genetic effects in complex traits such as AF.
Potential mechanism of genetic polymorphisms for the development of AF
Clinical studies have demonstrated that myocardial sleeves in PVs generate ectopic beats that play a substantial role in initiating and maintaining AF [14]. Electrical isolation of those sleeves is a cornerstone of most AF ablation procedures. PITX2c-deficient mice do not develop pulmonary myocardial sleeves [6]. Expression of PITX2 is remodeled during cardiac looping, becoming localized to the ventral portion of the developing ventricular chambers, while maintaining distinct left-sided atrial expression [20]. In addition, PITX2 might also trigger AF by altering the conduction properties of the cardiac conduction system. Wang et al. [21] have demonstrated that PITX2 haplo insufficiency predisposes individuals AF in electrically stimulated adult mice. Notably, ANK2, a gene known to play essential roles in in the localization and membrane stabilization of ion transporters and ion channels in disorders such as familial long QT syndrome, sinus node dysfunction, and AF, is also located near rs2200733 [22]. The ZFHX3 gene on chromosome 16q22 encodes a transcription factor that was originally identified as a regulator of alpha-fetoprotein expression [23]. ZFHX3 has been associated with Kawasaki disease, as well as malignancies such as prostate cancer [24]. However, in human cardiac and pulmonary tissue, its expression pattern and mechanism are not fully known. KCNN channels are also expressed in vascular endothelial cells, and suppression of KCNN3 expression in a mouse model has been associated with increased blood pressure [25]. In a rabbit burst-pacing model designed to simulate pulmonary venous ectopy, pharmacologic blockade of KCNN channels inhibited pacing-induced shortening of pulmonary venous and atrial action potential duration [26]. Shortening of the atrial action potential duration reduces the refractory period of atrial myocytes and promotes re-entry, an important mechanism for the development and maintenance of AF. Although the association of rs13376333 in KCNN3 with AF was not demonstrated in the present study, future research will be warranted to further investigate the association between them and elucidate the functional pathways.
The genetic polymorphisms and phenotypes of AF
Very few studies have evaluated the correlation of genetic variants, AF phenotypes, and response to AF therapies. Firouzi et al. [27] demonstrated increased dispersion of atrial refractoriness in patients with structurally normal hearts who carried the AA genotype of the connexin 40-promoter polymorphism. Husser et al. [28] found lower AF rates on surface ECG in individuals with the KCNE1 GG genotype. Here, we found the association of genetic polymorphisms with phenotypes of AF patients. Increased risk of AF from 1.04 to 3.9-fold has been observed in patients with prolonged PR interval, and prolonged PR interval is consistently one of the most significant phenotypes related to AF.[10] PR interval is a measure of atrial and atrio-ventricular nodal conduction, and can reflect left or right atrial hypertrophy or enlargement. A recent genome-wide association study of PR interval found links between PR interval and AF related genes such as CAV1-CAV2, NKX2-5, SOX5, SCN10A, SCN5A, and numerous other genes.[29] Our AF cohort shows a significant increase in PR interval and negative P wave terminal force in Lead V1 among patients with the rs2200733 risk allele. Negative P wave terminal force in Lead V1 is indicative of LA conduction impairment, or hypertrophy. Our results indicate that LA hypertrophy, or enlargement modulated by the 4q25 genetic variant, may increase the risk of AF development. Catheter ablation is one of the most effective treatments for patients with drug-refractory PAF or PeAF.[14] Many positive outcomes have emerged from the therapy, but adverse effects resulting from scar tissue formation and inability to restore the function of the treated myocardium, as well as complications related to the procedure remain as concerns. Therefore, a better way to predict recurrence of AF after catheter ablation is required. The exact mechanism of AF recurrence after catheter ablation remains elusive. However, multiple studies have shown an impaired clinical response to RFCA of AF in patients carrying the 4q25 rs2200733 variant allele. In a study by Shoemaker[30], a cohort of 378 patients who underwent catheter-based AF ablations showed 24% shorter recurrence-free time in patients with the rs2200733 risk allele compared to those that did not carry the risk allele. In our study, even though the association of rs2200733 with the recurrence of AF after RFCA was not statistically significant, a graded risk of AF recurrence was observed with increasing number of risk alleles at the 4q25 and ZFHX3 loci. The combination of rs2200733 and rs2106261 might serve as a risk factor for increased AF recurrence after catheter ablation. Our findings suggest that multi-marker allele combinations can be used as a clinical tool for the selection of patients for AF ablation. Further research will be warranted to determine whether risk stratification and patient management can be improved by the incorporation of genetic findings.
Study limitations
We analyzed the SNPs in a Korean population undergoing catheter ablation. Therefore, our findings cannot be generalized to all types of AF. When applying SNPs as predictors and targets of therapy for AF, possible racial differences in the association between SNPs and AF should be considered.
Conclusion
This study shows the common AF susceptibility SNP alleles of 4q25 and ZFHX3, are associated with the electroanatomical characteristics of LA and the clinical outcomes of catheter ablation in Korean patients with AF. These findings imply that gene variants may have a potential role for risk stratification of ablation therapy and post-ablation management.













